
- Southern blotting also called Southern hybridization is a hybridization technique which helps to detect specific gene sequences or DNA fragments within complicated mixture of nucleic acids.
- This technique is named after its inventor Edward N. Southern.
- This technique helps in the identification of a particular DNA fragment containing the gene of interest.
You may also want to see: Western blotting: Principle, Steps involved and Applications
Principle:
- This technique is based on the formation of hybrid DNA molecules when homologous, denatured DNAs from two different sources are mixed with each other under the appropriate conditions of ionic strength and temperature.
- This process of base pairing between complementary single-stranded DNA molecules is known as hybridization.
- In Southern blotting, DNA is first cut with restriction enzymes and the fragments are separated according to size by agarose gel electrophoresis.
- The gel is then placed in alkaline salt solution to denature the double stranded DNA.
- The resulting single stranded DNA is then transferred onto a nitrocellulose or nylon membrane either by capillary blotting or by electroblotting.
- After transfer, the strands on the membrane surface are immobilized by baking or UV-radiation.
- The membrane is then incubated with an appropriate radiolabeled probe (radiolabeled ss DNA or radiolabeled RNA) specific for the gene of interest.
- The probe hybridizes with the immobilized ssDNA on the membrane having the gene of interest.
- The position of the band containing these hybridized fragments is determined by autoradiography or any other method depending on the types of probes used.
- The size of the DNA fragments can be determined by comparing their relative size with the DNA bands of known lengths.
Steps involved:
1. Cleavage or digestion of DNA by restriction enzyme:
- DNA molecule having the gene of interest is digested into fragments using an appropriate restriction enzyme.
- The fragments obtained can be amplified by PCR.
2. Agarose gel electrophoresis:
- The sample having sufficient DNA fragments is loaded onto the wells on agarose gel.
- Electrophoresis is done after which the DNA fragments get separated on the gel according to their molecular weight.
3. Denaturation of ds DNA fragments:
- Agarose gel after electrophoresis is placed in alkaline salt solution (1.5 M NaCl and 0.5 M NaOH).
- This causes the denaturation of ds DNA fragments as it enhances the separation of DNA strands.
4. Capillary blotting or electroblotting:
- The ssDNA thus formed on agarose gel is then transferred to positively charged nitrocellulose or nylon membrane by capillary blotting.
- In case of polyacrylamide gel, capillary blotting is too slow and enough DNA might not be transferred. Hence, electroblotting is preferred in such case.
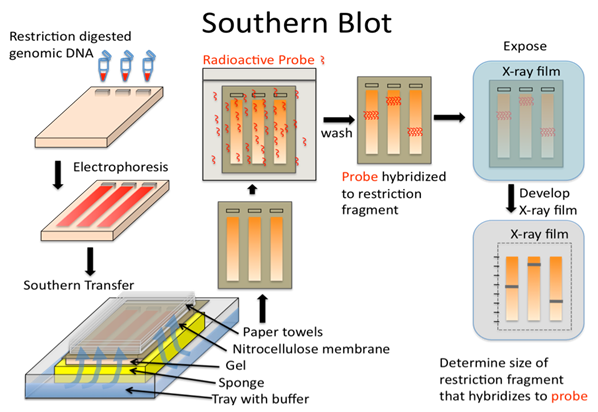
Image Credit:
https://bio.libretexts.org/Bookshelves/Genetics/Book%3A_Online_Open_Genetics_(Nickle_and_Barrette-Ng)/08%3A_Techniques_of_Molecular_Genetics/8.07%3A__DNA_Analysis-_Blotting_and_Hybridization
5. Baking and immobilization:
- Once the ssDNA is transferred, the membrane is slightly heated in an autoclave to fix the membrane, the process called baking.
- After baking, casein or Bovine serum albumin (BSA) is added to the membrane which masks off all the binding sites of the membrane.
6. Hybridization with labeled probe:
- The membrane is then incubated with a labeled probe which contains the complementary sequences to the gene of interest on the membrane.
- The probe can be labeled with any radioactive substance or dyes with fluorescence or chromogenic property.
- The base pairing between the probe and ssDNA on the membrane forms a hybrid DNA molecule by the process called hybridization.
7. Detection and visualization by autoradiography:
- The hybridized fragments on the membrane can be detected by autoradiography.
- If a fluorescent or a chromogenic dye is used, the hybridized regions can be visualized on X-ray film or as colored spots on the membrane.
Applications:
- It is used to detect and determine the size of specific DNA fragment in the given mixture of genomes.
- It is applied in the field of gene mapping and evolution.
- It can be used in diagnostic study of genetic diseases.
- The technique can be used for DNA analysis to detect point mutations and other structural rearrangements in the DNA sequences.
- Used for paternity testing along with the identification of criminals and victims.
- It is used in DNA fingerprinting and restriction fragment length polymorphism.
- It can also be used in the identification of the causative agent of any infectious disease.