
- Polymerase chain reaction (widely known as its abbreviated form PCR), is used for the amplification of specific segments or sequences of DNA or RNA.
- PCR technique was developed by Kary Mullis in 1983.
- It is a molecular cum enzymatic method which is carried out entirely in vitro.
Principle:
- PCR procedure is highly sensitive, simple and inexpensive that helps in characterization, analysis and synthesis of specific fragments of DNA or RNA from virtually any living organisms.
- PCR uses the enzyme DNA polymerase that directs the synthesis of DNA from deoxy-ribonucleotide substrates on a single-stranded DNA template.
- Two synthetic, single-strand oligonucleotides (15-30 bases long) are synthesized which are complementary to the sequences on the opposite strands of the target DNA at positions defining the ends of the segment to be amplified.
- DNA polymerase adds nucleotides to the 3’ end of these single-strand oligonucleotides called primers, when they are annealed to a longer template DNA.
- This reaction generates double-stranded DNA over the region of interest on both of the strands of DNA, which is the first cycle of PCR.
Steps in PCR:
- One cycle of PCR consists of the following three basic steps:
- Denaturation of DNA to be amplified:
- Isolated DNA, containing the segment to be amplified is heated at 920C to 960C for about 2 minutes.
- After heating, two strands of DNA separate or melt down to form single stranded DNAs.
- Annealing of primers to the template DNA:
- Annealing of primers to each strand is carried out after briefly cooling them down (450C to 550C).
- Extension or polymerization of the annealed primers:
- It is carried out at 720
- At this point, DNA polymerase and four deoxy-ribonucleoside triphosphates (also called deoxy-ribonucleotide substrates) are added.
- Heat stable DNA polymerase such as the Taq polymerase (derived from Thermus aquaticus living in hot springs) is used which is not denatured by the heating steps.
- DNA polymerase adds deoxy-ribonucleotides (dNTPs), complementary to the template strands at 3’ends of respective primers. i. e. the primed DNA segments are replicated selectively.
- Denaturation of DNA to be amplified:
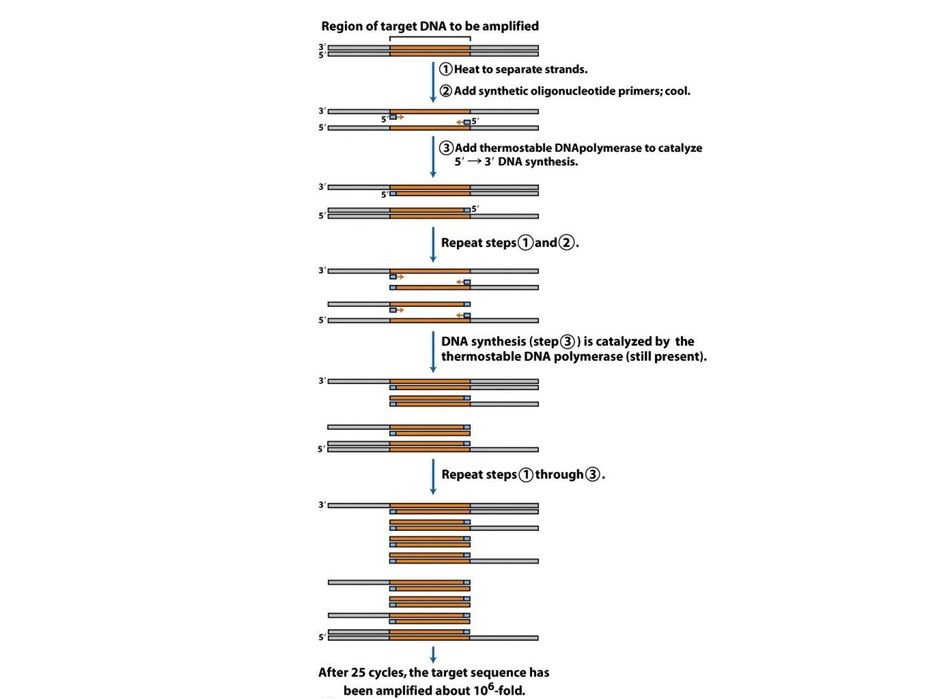
- After the completion of each cycle, two copies of DNA samples are produced. The doubling of number of DNA strands corresponding to target sequences can be estimated by amplification number associated with each cycle using the formula;
Amplification number =2n, where n=number of PCR cycles.
- The cycle of heating, cooling and replication is repeated 25-30 times over a few hours in an automated process in a thermocycler, amplifying the DNA segment between the primers until it can be readily analyzed or cloned.
Applications of PCR:
- Molecular archaeology and molecular paleontology: To clone DNA fragments from the mummified remains of humans and extinct animals
- Epidemiology: Epidemiologists can use PCR-enhanced DNA or RNA samples from human remains to trace the evolution of human pathogenic viruses.
- Forensic science: DNA finger printing, paternity testing and criminal identification
- Diagnosis of infection or disorder:
- Molecular detection and identification of viral infections before they cause symptoms
- For the prenatal diagnosis of a wide range of array of genetic diseases
- Evolutionary biology: To track down and trace the evolutionary history of organisms
- For the study of mutation
- Whole genome sequencing: Mapping of expressed gene sequence
- Gene cloning and gene expression
- Drug discovery and vaccine production by recombinant DNA technology
- Human genome project