
- Regulation of gene expression involves two steps; transcription and translation to form a complete protein or enzyme.
- Some enzymes are required under certain conditions but not all times. Synthesis of such enzymes continuously all the time, is wastage of energy.
- In order to minimize this energy loss, some control mechanisms are required to control transcription and translation.
- These mechanisms allow transcription and translation to occur only when the enzymes are required and block the same when the enzymes are not required.
- Such control over transcription and translation as per the requirement of certain protein is called regulation of gene expression.
- One best example of regulation of gene expression is Lac Operon in E. coli.
Lac Operon in E. coli:

https://www.toppr.com/ask/en-bd/question/which-is-not-component-of-lac-operon/
- An operon is a functional unit of DNA that contains a cluster of genes which are controlled by a single promoter. Such genes present in the operon are either expressed together for a particular protein or not at all.
- ‘Lac’ in Lac Operon stands for lactose. Lac Operon is an example of control of transcription.
- It controls the transcription of genes that code for the enzymes required for the transport and metabolism of lactose in many enteric bacteria including coli.
- Lac Operon consists of two types of genes; structural genes and regulatory genes.
- Structural genes of Lac Operon are lac Z, Lac Y and Lac A which encode for respective enzymes required for the transport and metabolism of lactose.
- lac Z: It encodes the enzyme β-galactosidase which hydrolyzes lactose to form glucose and galactose and also converts lactose into its isomeric form, allolactose.
- lac Y: It encodes the enzyme β-galactoside permease which is a transporter protein and facilitates the entry of lactose into the cell.
- lac A: It encodes the enzyme β-galactoside transacetylase. This enzyme transfers an acetyl group from acetyl-CoA to toxic β-galactosides, glucosides and lactosides and removes them out of the cell.
- Regulatory genes include a repressor gene (Lac I) along with Promoter (P) and Operator (O) genes.
- Regulatory genes regulate the transcription of structural genes.
- lac I: It encodes repressor protein that binds to the operator sequence.
- Promoter gene: It serves as the binding site for RNA polymerase.
- Operator gene: It serves as the binding site for repressor protein.
Mechanism of regulation of Lac Operon:
- Lac Operon is regulated by the following three mechanisms:
- Repression of Lac mRNA transcription
- Induction of Lac mRNA transcription
- Catabolic repression: (In the presence of glucose and lactose)
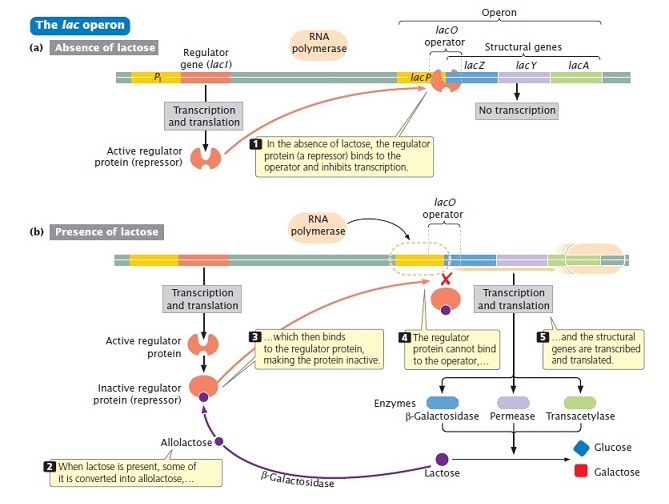
Repression of Lac mRNA transcription (In the absence of lactose):
- RNA polymerase must bind to promoter gene in order to transcribe structural genes.
- Repressor gene (lac I) encodes repressor protein that binds to operator gene.
- There are three regions in operator gene. O1, O2 and O3.
- Repressor protein generally binds to region O1 of Operator but sometimes can bind to secondary region O2 which is near lac Z and region O3 which is upstream to promoter.
- When repressor protein binds to operator, RNA polymerase cannot bind to promoter. It is because binding of repressor to operator overlaps promoter and prevents RNA polymerase from binding to promoter.
- No binding of RNA polymerase to promoter, stops the transcription of structural genes. This occurs in the absence of lactose in the cell.
- Some coli even in their repressed state have minimal level of enzymes, β- galactosidase and β-galacotoside permease which are essential for operon regulation in the immediate supply or lactose afterwards.
https://microbiochem.weebly.com/603uiiilacoperon
Induction of Lac mRNA transcription (In the presence of lactose):
- When lactose is present in the cell, it binds to repressor protein causing its conformational change. This change dissociates repressor protein form the operator.
- Lactose appears to serve as an inducer here but actually it is its isomeric form, allolactose, which acts as a real inducer.
- β-galactosidase converts lactose into allolactose that binds to repressor protein and inactivates it.
- Inactivation of repressor protein reduces its affinity towards the operator. This leads to the binding of RNA polymerase to the promoter region.
- Finally, transcription of structural genes occurs and the enzymes β-galactosidase, β-galactoside permease and β-galactoside transacetylase are synthesized responsible for the metabolism of lactose.
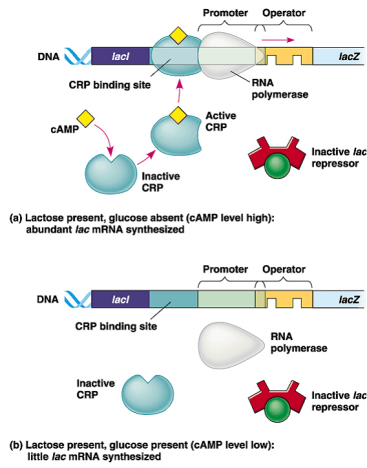
Catabolite repression or positive control (In the presence of glucose and lactose):
https://www.ourbiochemistry.com/knowledge-base/lac-operon-regulation-of-gene-expression-in-prokaryotes-lecture-2/
- When glucose is present in the cell, metabolism of lactose doesn’t occur as glucose suppresses the transcription of lactose metabolizing enzymes.
- It is regulated by catabolite activator protein (CAP) encoded by CAP gene located elsewhere in the chromosome.
- The binding of CAP near promoter upstream to Lac I facilitates the binding of RNA polymerase to promoter more tightly. This tight binding accelerates the rate of transcription by 20 folds.
- CAP initially is inactive which requires cyclic adenosine monophosphate (cAMP) for activation. cAMP, being a secondary messenger, binds to CAP only in the absence of glucose.
- When glucose level increases in the cell, it inhibits the synthesis of cAMP. High glucose level also transports cAMP out of the cell decreasing its concentration in the cell.
- Hence, when glucose level increases, cAMP level decreases and CAP is not activated, finally inhibiting transcription of Lac genes for lactose metabolism.
Feedback regulation of lac operon:
- Active transport of lactose (inducer) into the cell by β-galactoside permease increases its concentration thus resulting in increased induction of lac operon.
- Induction of lac operon is enhanced by transgalactosylation of lactose by β-galactosidase producing allolactose.
- Increased concentration of glucose due to the hydrolysis of lactose and allolactose by β -galactosidase results in catabolite repression of lac operon.